|
Khuyen and Michael successfully defended their thesis proposals and advanced to Masters candidacy recently. Congratulations and best wishes to them on finishing up their theses in the coming term!
0 Comments
Congratulations to Anais Aoki on a splendid Masters thesis proposal defense, which she passed with flying colors and officially advanced to Masters candidacy. We can't wait to hear more about her ongoing research on predictive models for conservation genomics!
The Sethuraman Lab invites applications from potential PhD and Masters students for Fall 2023 - we are looking for scholars with a background and research interest in (1) population genetics/genomics, (2) bioinformatics/computational biology, (3) evolutionary biology, (4) conservation, (5) computer science/applied mathematics, (6) machine learning to join the lab through several graduate programs at San Diego State University:
PhD programs 1) Joint Doctoral Program in Evolutionary Biology (with University of California Riverside) 2) Joint Doctoral Program in Computational Sciences (with University of California Irvine) Masters programs 1) Masters in Evolutionary Biology 2) Masters in Bioinformatics and Medical Informatics This recruitment season, we are specifically looking for PhD students interested in the following funded projects: 1) Developing new statistical methods/software for estimating evolutionary history - for examples of methods that we have developed, see: PPP, InRelate, IMa2p, IMGui, MigSelect, and MULTICLUST. 2) Understanding human evolutionary history, specifically in the face of gene flow from archaic ghost populations. We are interested in understanding/estimating signatures of selective sweeps, linked selection, adaptive/maladaptive introgression, load, and demographic history across diverse human populations. Potential Masters students are welcome to write to me with their broader interests and we can discuss more about potential projects. Please write to Dr. Sethuraman (asethuraman@sdsu.edu) with a copy of your CV/Resume, a brief statement of interest, and let’s chat more. We are committed to recruiting a diverse group of scientists to join my lab group – so we highly encourage folks who identify as a part of historically underrepresented groups to apply. This includes (non-exhaustively) people of color, international students, Veterans of armed forces, student-parents/caregivers, first-generation degree holders, the LGBTQIA+ family, and folks with medical conditions and disabilities. PI Sethuraman was at the PEQG 2022 meeting in Pacific Grove, CA to present the lab's recent work describing the multinomial clustering method for estimating population genomic structure from polyploid, multi-allelic genomic data. Watch for the preprint describing MULTICLUST! Meanwhile, the tool is accessible via GitHub here.
We are so proud of our lab's three graduates from the Class of 2022 - Robert Reyes, Walker Welch, and MJ Lee! Congratulations and we can't wait to see what amazing things you will go on to do!
Our manuscript that describes the genome of a novel Sediminibacterium discovered in association with two separate cyanobacterial isolates from freshwater locales in Southern California has now been accepted at G3! All the analyses for this manuscript were performed by our super talented NSF REU scholars (including the Sethuraman Lab's very own Andrew Zhang), and the entire paper was written by all of us in a conference room, masked, with everyone contributing sentences - the most unique experience of all! Read the preprint here, while we await the print version!
Our manuscript detailing the changes in microbial diversity and community composition in response to chronic nitrogen input using a long-term experiment has now been published in Applied Soil Ecology! We discovered that soil inorganic nitrogen and pH were prominent predictors of microbial abundance - all work led by Tim Grant from CSUSM. Read more here. 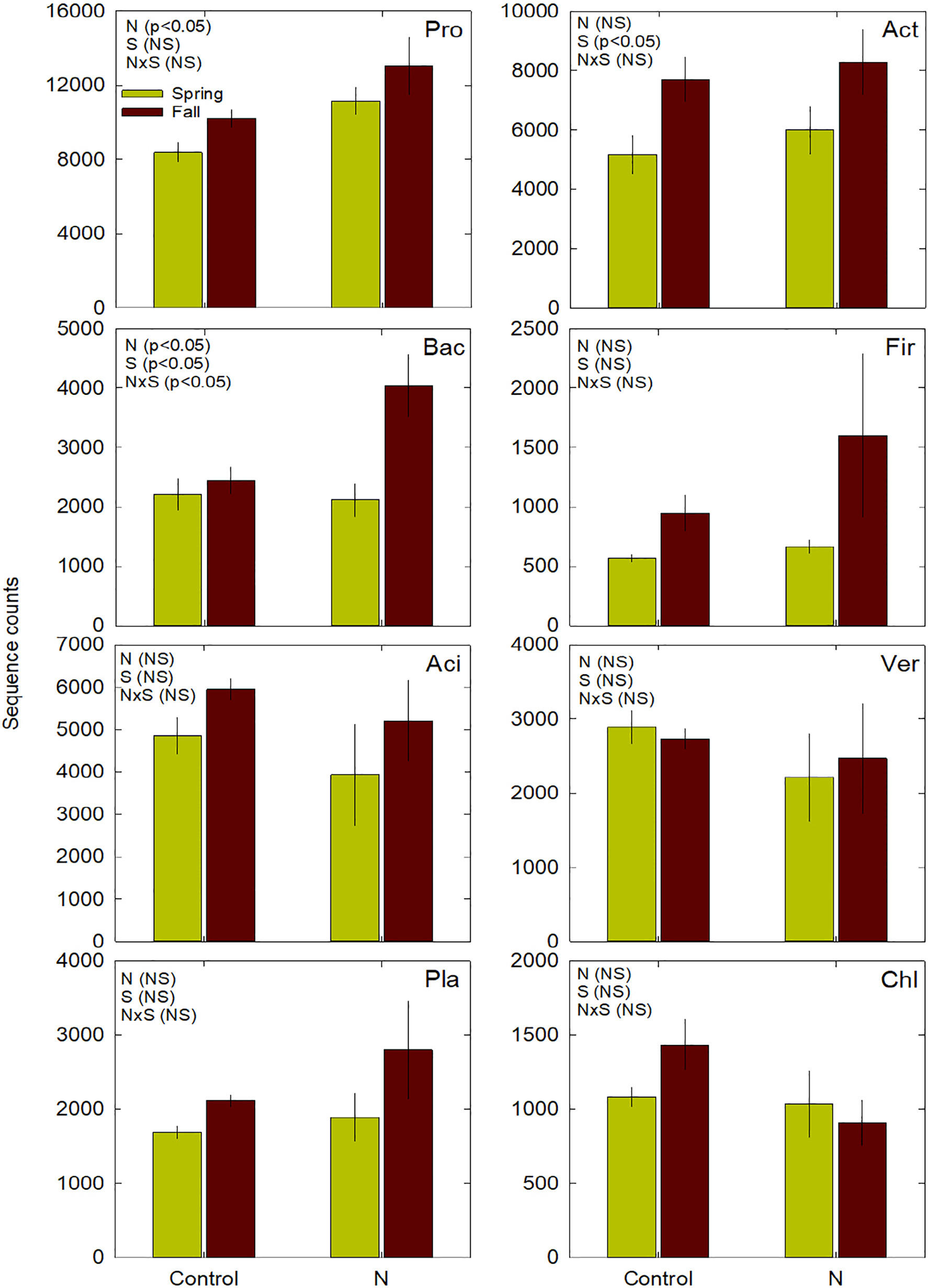 Mean (±SE; n = 4 plots/treatment) sequence counts for Proteobacteria (Pro), Actinobacteria (Act), Bacteroidetes (Bac), Firmicutes (Fir), Acidobacteria (Aci), Verrucomicrobia (Ver), Planctomycetes (Pla), and Chloroflexi (Chl) bacterial phyla in coastal sage scrub (CSS) exposed to ambient (control) and elevated N (N) during the spring (green) and fall (brown). Also shown are results of a two-way ANOVA or permutation test with 1 factor and 12 error degrees of freedom (p-value only) for effects of treatment (N), season (S), and their interaction (N × S). NS = not significant (p > 0.05). Several of our Sethuraman Lab members presented their research at the SDSU Student Research Symposium on Friday, 3/4/2022 at Montezuma Hall. Congrats to everyone on presenting all the fun science!
 The first chapter from Alicia's Masters thesis describing the genome of the parasitoid wasp Dinocampus coccinellae and comparative phylogenomics of hymenopterans has now been published in G3! Our study determined extensive duplications, increased rates of gene gain/loss along the D. coccinellae lineage, along with re-affirming multiple independent origins of eusociality and thelytokous parthenogeny among hymenoptera. Read the full paper here. Congratulations to all Sethuraman Lab students who were instrumental in this publication! Also, this stunning image showing a larval D. coccinellae egressing from an adult Coccinella septempunctata captured by Roxane Saisho is now featured on the cover of the March 2022 issue of G3! Check out the entire issue here.
Dr. Sethuraman was recently featured on the latest edition of UC Merced's RadioBio podcast where he talks about his work on ghost populations, and population genomics of biological control. Give it a listen here!
|
Arun Sethuraman
|











 RSS Feed
RSS Feed