|
The Sethuraman Lab recently presented results from numerous projects at the SDSU Student Symposium (S3) - Raya on her Undergraduate capstone work, developing a new tool called speciel for developing SNP arrays to identify trout species, Tamsen on her PhD work, utilizing Ks distributions to estimate and distinguish timing of allo- and auto-polyploidization events, and Priyanshi on her MS thesis on quantifying telomere length variation in diverse human populations, and the development of a deep learning framework to estimate tumor status. Priyanshi was awarded the Provost's Award for the Sciences, and Tamsen was awarded the Excellence in Graduate Thesis Award! Congratulations to both of them!
0 Comments
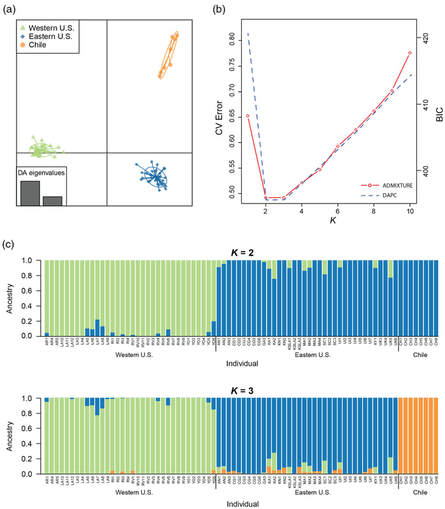
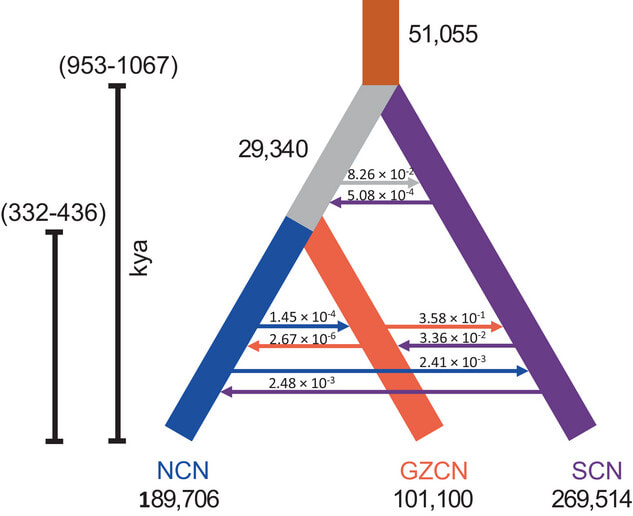
Our lab's work describing range-wide population genomics of the convergent lady beetle, Hippodamia convergens is now out in Evolutionary Applications! We describe the efficacy of using genotyping by sequencing techniques to assess the efficacy of augmentative biological control in the species, showing significant degree of population structure, no contemporary gene flow between Eastern US, Western US, and Chilean populations. Read our work here.
New manuscript on genome-wide adaptations for invasion success in Sesamia inferens now out!1/24/2024 Our new manuscript describing genome-wide adaptations to invasion in the pink rice borer, Sesamia inferens is now out in Insects. We use a combination of evolutionary population genomics and comparative analyses to estimate strong selection at insect cuticle glycine-rich cuticular protein genes which are associated with enhanced desiccation adaptability, and at the histone-lysine-N-methyltransferase gene associated with range expansion and local adaptation. Read here! Congrats to Scott Monahan and Ryan Buck on co-authoring this massive work!
Dr. Sethuraman was recently elected to the Board of Directors of the Genetics Society of America, where he will serve from 2024-2026 alongside fellow new electees, Dr. Jason Stajich (UC Riverside) and Dr. Eyleen O'Rourke (University of Virginia). We look forward to continued involvement and leadership in the GSA!
Congratulations to our lab's newest Masters candidate, Priyanshi on successfully defending her thesis proposal on "Predictive modeling of tumor status in The Cancer Genome Atlas data". Watch this space for updates and new discoveries from her research.
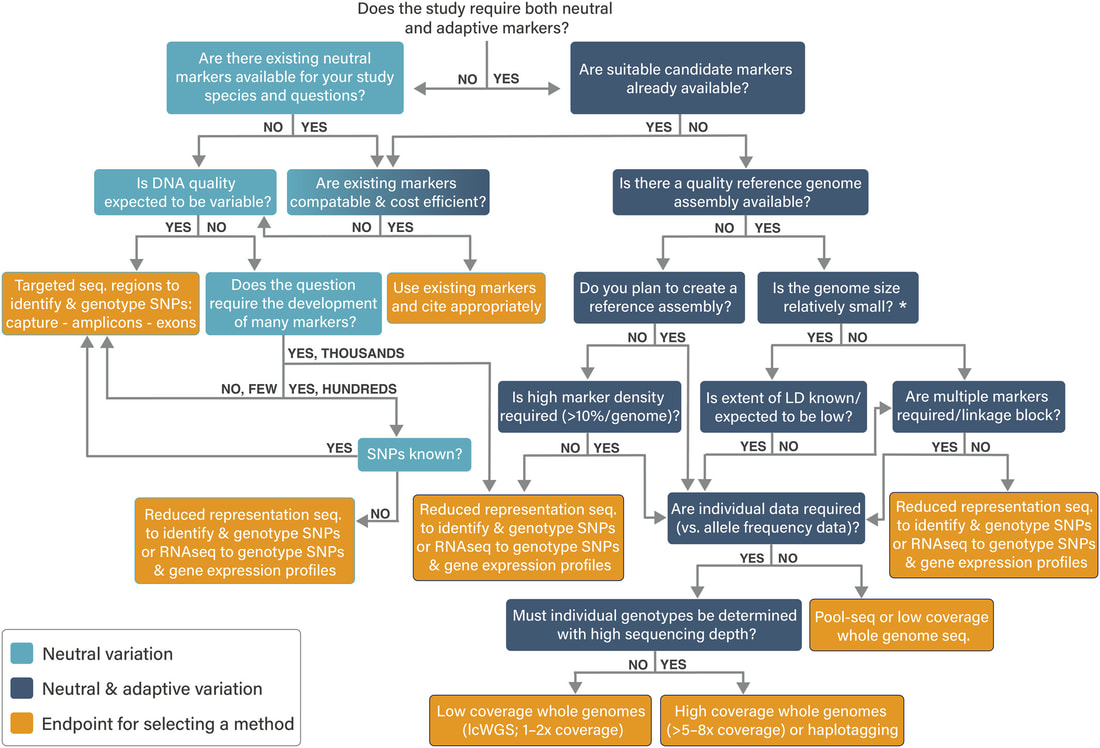
New review on population genomics in conservation published in Molecular Ecology Resources11/16/2023 Our new invited review on using population genomics to inform conservation is now published in Molecular Ecology Resources! We describe a host of resources, review current methods/methodologies, technologies, software, and best practices for their use. We also provide a series of tutorials/resources for budding conservation genomicists. Do check out the manuscript here. Congratulations to Lauren and the entire ConGen2022 team for this massive work!
PhD student Alex recently received the 2024 California State University Agricultural Research Institute's Nextgen Fellowship, and she joins a cohort of students across the CSU system to study plant resilience in the face of climate change! This fellowship will fund a multi-year field + greenhouse experiment to understand drought tolerance in US-based hop cultivars. Congratulations, Alex!
Tamsen, Alex, Trevor and Priyanshi recently received the Genetics Society of America's Presidential Membership Initiative Award! Congratulations, everyone!
Our lab recently received the 2023 CSUBIOTECH Faculty-Student Collaborative Research: Development Grant, which will support our Masters student, Trevor Mugoya for two years, along with one undergraduate student. Congrats everyone!
PhD student Alex recently presented her research on hops at the Women in STEM event at the SDSU campus on October 23 - we're en route sequencing a hundred hop genomes to study their evolutionary history. We can't wait to share updates on this project!
|
Arun Sethuraman
|



 RSS Feed
RSS Feed